AP22686112 – Investigation of somatic mutations based on single-cell RNA using machine learning methods in patients with peripheral artery disease
The main goal of this study is to create an inclusive pipeline for detecting somatic mutations in patients with peripheral artery disease (PA) using tools such as Gemini, Cosmic and Monocle, as well as various machine learning methods. The goal of the first year is to review and write an overview article about precomputation tools and gene expression clustering tools.
Relevance: Peripheral arterial disease (PD), as part of cardiovascular diseases (CVD), remains one of the leading causes of disability and mortality. Modern diagnostic methods do not always make it possible to identify individual genetic predispositions. The development of an effective pipeline for detecting somatic mutations using AI and genetic tools will improve diagnostic accuracy, speed up the choice of therapy and reduce the burden on the healthcare system.
Scientific supervisor: Kunikeyev Aydin
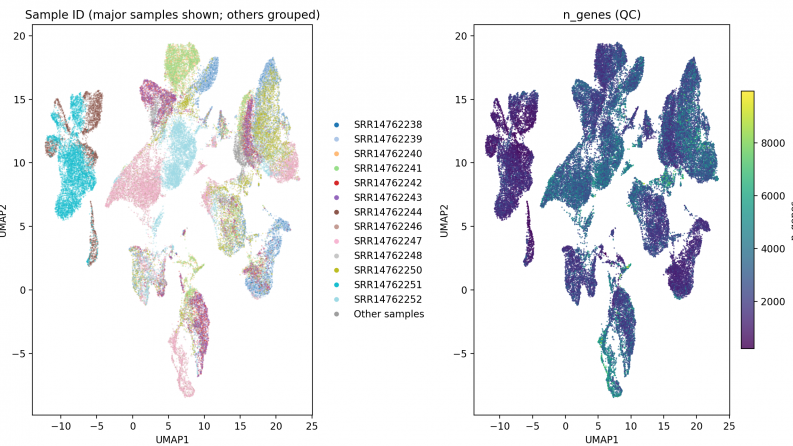
Expected and achieved results:A clustering analysis was performed on the processed scRNA-seq dataset to identify cell types. A reproducible pipeline based on PRJNA736095 data was implemented, covering all stages from raw reads to final cell annotation. Quality control, normalization, highly variable gene selection, PCA, UMAP, and Leiden clustering were conducted. As a result, major immune and stromal cell populations (T, NK, B/plasma, myeloid, dendritic, endothelial, etc.) were reliably identified. Marker genes were calculated for each cluster, enabling biological interpretation. Consistent UMAP visualizations with unified color schemes ensured comparability across samples. The obtained results formed the basis for preparing a scientific publication and a doctoral dissertation.
